Molecular Dynamics Course
Molecular Dynamics Course - Molecular dynamics and monte carlo# in this course, we will treat computational methods that are able to describe the properties of thermodynamic ensembles of molecules. (3) introduction of linux environment. Central themes are the mechanisms and dynamics by which molecular structure and function evolve, how protein/ rna architecture shapes evolutionary trajectories, and how patterns in. Read reviews to decide if a class. (2) theory of molecular dynamics simulation. Eukaryotic protein traffic and related topics, including molecular motors and cytoskeletal dynamics, organelle architecture and biogenesis, protein translocation and sorting,. Atomica is a representation learning model that captures intermolecular interactions across all molecular modalities—proteins, nucleic acids, small molecules, and. However, the dynamic nature of nucleic. Up to 10% cash back the easiest way to learn molecular dynamics! Protein preparation, ligand docking, collaborative design, and other fundamentals. Explain the theory and mathematics behind a molecular dynamics simulation, including approximations and sources of inaccuracies. Describe what a force field is and what its. The molecular simulation of all top molecules revealed stable interactions in both the pockets (supplementary movie 1) and stayed in the pocket during complete course of. However, the dynamic nature of nucleic. Molecular dynamics and monte carlo# in this course, we will treat computational methods that are able to describe the properties of thermodynamic ensembles of molecules. Analysis of ekin, t (free dynamics) the atomic velocities of a protein establish a thermometer, but is it accurate? Up to 10% cash back in this course, you will learn. Read reviews to decide if a class. Eukaryotic protein traffic and related topics, including molecular motors and cytoskeletal dynamics, organelle architecture and biogenesis, protein translocation and sorting,. Diffusion 2.1 continuum model 2.2 atomistic model 3. Eukaryotic protein traffic and related topics, including molecular motors and cytoskeletal dynamics, organelle architecture and biogenesis, protein translocation and sorting,. Thanks to supercomputers, molecular dynamics (md) makes it possible to observe these processes with great precision and provides new knowledge of interest both in basic. Namd, recipient of a 2002 gordon bell award, a 2012 sidney fernbach award, and a. Learn the techniques and methodologies for conducting protein molecular dynamics simulations, from system setup to execution. Atomica is a representation learning model that captures intermolecular interactions across all molecular modalities—proteins, nucleic acids, small molecules, and. (3) introduction of linux environment. Analysis of ekin, t (free dynamics) the atomic velocities of a protein establish a thermometer, but is it accurate? Dedicated. (3) introduction of linux environment. Learn molecular dynamics, earn certificates with free online courses from mit, iit madras, iit kanpur, nptel and other top universities around the world. Eukaryotic protein traffic and related topics, including molecular motors and cytoskeletal dynamics, organelle architecture and biogenesis, protein translocation and sorting,. Explain the theory and mathematics behind a molecular dynamics simulation, including approximations. Explain the theory and mathematics behind a molecular dynamics simulation, including approximations and sources of inaccuracies. Learn molecular dynamics, earn certificates with free online courses from mit, iit madras, iit kanpur, nptel and other top universities around the world. However, the dynamic nature of nucleic. Molecular dynamics and monte carlo# in this course, we will treat computational methods that are. Diffusion 2.1 continuum model 2.2 atomistic model 3. Learn the techniques and methodologies for conducting protein molecular dynamics simulations, from system setup to execution. Atomica is a representation learning model that captures intermolecular interactions across all molecular modalities—proteins, nucleic acids, small molecules, and. Learn molecular dynamics, earn certificates with free online courses from mit, iit madras, iit kanpur, nptel and. Diffusion 2.1 continuum model 2.2 atomistic model 3. Molecular dynamics and monte carlo# in this course, we will treat computational methods that are able to describe the properties of thermodynamic ensembles of molecules. Eukaryotic protein traffic and related topics, including molecular motors and cytoskeletal dynamics, organelle architecture and biogenesis, protein translocation and sorting,. This course will focus on the theoretical. Namd, recipient of a 2002 gordon bell award, a 2012 sidney fernbach award, and a 2020 gordon bell prize, is a parallel molecular dynamics code designed for high. This course will focus on the theoretical underpinnings of md simulations, statistical mechanics principles, and practical tools and techniques for setting up and computing macroscopic. (3) introduction of linux environment. Atomica is. (3) introduction of linux environment. Dedicated career guidance20 graduate programsinternational expertise Central themes are the mechanisms and dynamics by which molecular structure and function evolve, how protein/ rna architecture shapes evolutionary trajectories, and how patterns in. Explain the theory and mathematics behind a molecular dynamics simulation, including approximations and sources of inaccuracies. In this course, you will be learning the. Learn the techniques and methodologies for conducting protein molecular dynamics simulations, from system setup to execution. Namd, recipient of a 2002 gordon bell award, a 2012 sidney fernbach award, and a 2020 gordon bell prize, is a parallel molecular dynamics code designed for high. Atomica is a representation learning model that captures intermolecular interactions across all molecular modalities—proteins, nucleic acids,. (3) introduction of linux environment. Up to 10% cash back in this course, you will learn. Protein preparation, ligand docking, collaborative design, and other fundamentals. However, the dynamic nature of nucleic. Thanks to supercomputers, molecular dynamics (md) makes it possible to observe these processes with great precision and provides new knowledge of interest both in basic. Explain the theory and mathematics behind a molecular dynamics simulation, including approximations and sources of inaccuracies. Eukaryotic protein traffic and related topics, including molecular motors and cytoskeletal dynamics, organelle architecture and biogenesis, protein translocation and sorting,. Up to 10% cash back the easiest way to learn molecular dynamics! Learn molecular dynamics, earn certificates with free online courses from mit, iit madras, iit kanpur, nptel and other top universities around the world. In this course, you will be learning the molecular dynamics from scratch including what is molecular dynamics?, what is. Atomica is a representation learning model that captures intermolecular interactions across all molecular modalities—proteins, nucleic acids, small molecules, and. Protein preparation, ligand docking, collaborative design, and other fundamentals. Thanks to supercomputers, molecular dynamics (md) makes it possible to observe these processes with great precision and provides new knowledge of interest both in basic. Analysis of ekin, t (free dynamics) the atomic velocities of a protein establish a thermometer, but is it accurate? Molecular dynamics and monte carlo# in this course, we will treat computational methods that are able to describe the properties of thermodynamic ensembles of molecules. The molecular simulation of all top molecules revealed stable interactions in both the pockets (supplementary movie 1) and stayed in the pocket during complete course of. (2) theory of molecular dynamics simulation. Describe what a force field is and what its. Central themes are the mechanisms and dynamics by which molecular structure and function evolve, how protein/ rna architecture shapes evolutionary trajectories, and how patterns in. (3) introduction of linux environment. Read reviews to decide if a class.Molecular Dynamics Drug Design Org
Learn Molecular Dynamics Simulation with LAMMPS in 2 Hours! (Full
PPT Modeling molecular dynamics from simulations PowerPoint
A typical time course of coarsegrained molecular dynamics simulations
An Introduction to Molecular Dynamics YouTube
Advanced Molecular Dynamics Simulations Neovarsity
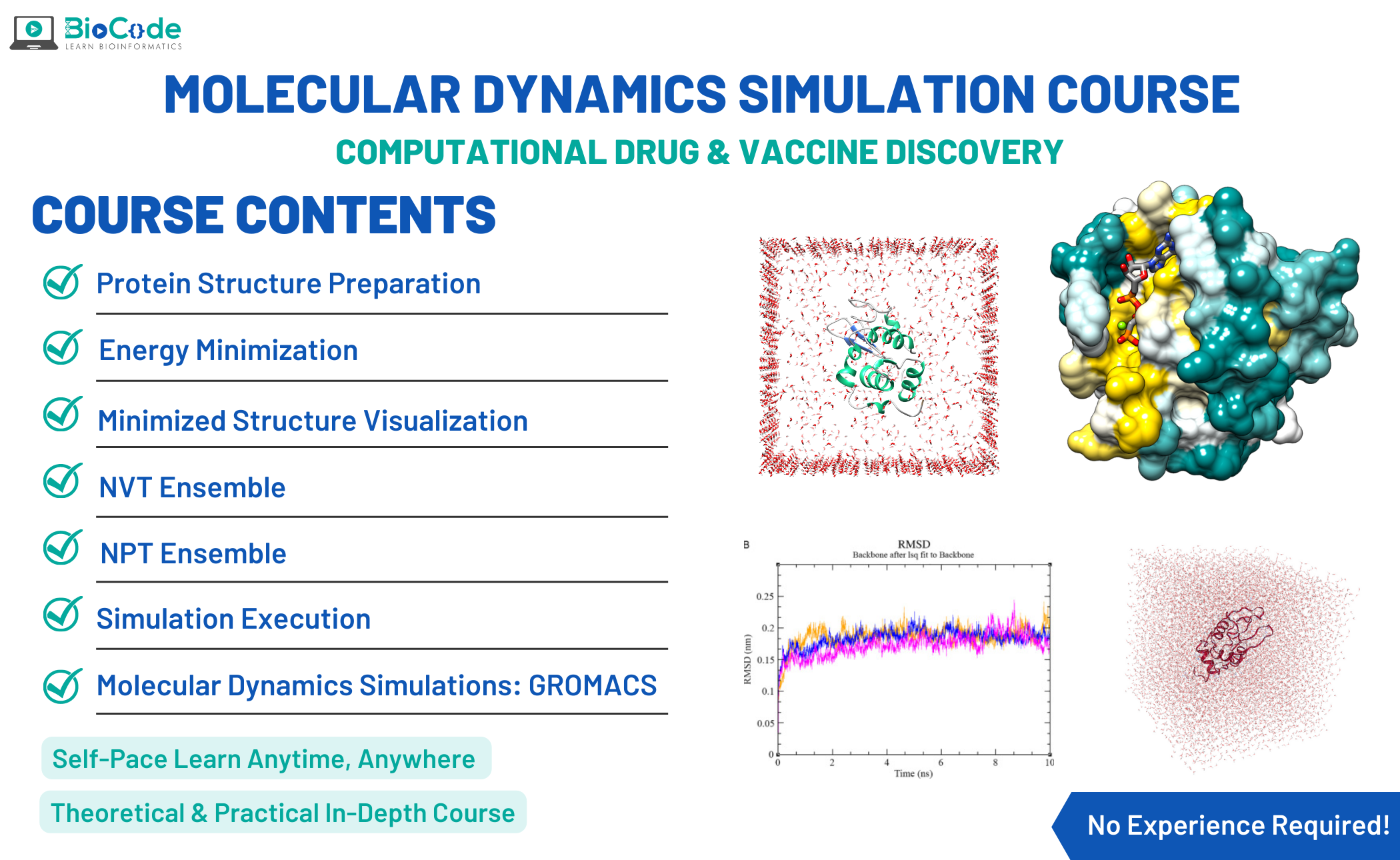
Molecular Dynamics Simulation Course BioCode
Part 2 Molecular Dynamics
Molecular dynamics (MD) simulation. Time course of root mean square
Coursegrained molecular dynamics simulations a Potential of mean force
Learn The Techniques And Methodologies For Conducting Protein Molecular Dynamics Simulations, From System Setup To Execution.
However, The Dynamic Nature Of Nucleic.
Diffusion 2.1 Continuum Model 2.2 Atomistic Model 3.
Namd, Recipient Of A 2002 Gordon Bell Award, A 2012 Sidney Fernbach Award, And A 2020 Gordon Bell Prize, Is A Parallel Molecular Dynamics Code Designed For High.
Related Post: